2024
Utsab GhimireG, Eleni Pliakoni, Jeffrey Brecht, and Liu T. (2024) Unraveling the molecular mechanisms of 1-Methylcyclopropene (1-MCP) retards postharvest senescence in broccoli (Brassica oleracea). Postharvest Biology and Technology.
Garcia EG., Koh J., Wu X., Sarkhosh A, and Liu T. (2024) Tissue-specific proteome profile analysis reveals regulatory and stress-responsive networks in passion fruit (Passiflora edulis) during storage. Scientific Report.
Yogesh Ahlawatf, Song Li, Eleni D. Pliakoni, Jeffrey Brecht, Liu T. (2024) Tandem Mass Tag-based proteome analysis of postharvest senescence in broccoli (Brassica oleracea). Vegetable Research.
Kathryn ChaseG, Catherine Belisle, Yogesh Ahlawatf, Fahong Yu, Gérman Sandoya, Kevin Begcy, and Liu T.(2024) Examining preharvest genetic and morphological factors contributing to lettuce (Lactuca sativa L.) shelf life during natural senescence in broccoli (Brassica oleracea). Scientific Report.
Garcia EG., Koh J., Wu X., Sarkhosh A, and Liu T. (2024) Tissue-specific proteome profile analysis reveals regulatory and stress-responsive networks in passion fruit (Passiflora edulis) during storage. Scientific Report. Accepted.
Habibi,F., Boakye, D., Chang,Yu g, Casorzo,G. g, Hallman,L., Madison, M., Clavijo-Herrera,J g., Sarkhosh, A., Liu, T. (2024) Molecular mechanisms underlying postharvest physiology and metabolism of fruit and vegetables through multi-omics technologies. Scientia Horticulturae. 324, 112562. https://doi.org/10.1016/j.scienta.2023.112562
Liu T, Talavera-Rauh F. (2024) Regulation of class I TCP transcription factors by REVOLUTA and KANADI genes that is involved in leaf development in Arabidopsis. Under review.
2023
Utsab Ghimire, Eleni Pliakoni, Fahong Yu, Jeffrey K. Brecht, Liu T (2023) Identifying genes regulated during natural, on-plant senescence in broccoli (Brassica oleracea) in contrast to postharvest senescence. Postharvest Biology and Technology. 1206, 112535. https://doi.org/10.1016/j.postharvbio.2023.112535
Xiaolei Guo, Chiwah Tseung, Alina Zare, Liu T. (2023) Hyperspectral Image Analysis for the Evaluation of Chilling Injury in Avocado Fruit during Cold Storage. Postharvest Biology and Technology. https://doi.org/10.1016/j.postharvbio.2023.112548.
Habibi F, Liu T, Muhammad S, Schaffer B, Sarkhosh A. (2023) Physiological, Biochemical, and Molecular Responses of Fruit Trees to Rootzone Hypoxia. Environmental Experimental Botany. 206. February, 2023.
https://doi.org/10.1016/j.envexpbot.2022.105179
2022
Chang Y, Yogesh Ahlawat, Ali Sarkhosh, and Liu T. (2022) Spatiotemporal transcriptome mapping of muscadine berry development and ripening. Frontier in Plant Sciences. https://doi.org/10.3389/fpls.2022.949383
Liu T. Albornoz K., Deltsidis A., Beckles, D. (2022) Editoral: Postharvest Ripening, Senescence, and Technology. Frontier in Genetics. https://www.frontiersin.org/articles/10.3389/fgene.2022.920584/full
X Guo g, Y Ahlawat &, Liu T. A Zare, (2022) Evaluation of Postharvest Senescence in Broccoli Via Hyperspectral Imaging. Plant Phenomics.
https://spj.sciencemag.org/journals/plantphenomics/2022/9761095/
Fariborz Habibi, Liu T , Kevin Folta, Ali Sarkhosh. (2022) Physiological, Biochemical, and Molecular Aspects of Grafting in Fruit Trees. Horticulture Research. https://academic.oup.com/hr/advance-article/doi/10.1093/hr/uhac032/6532224
Yogesh Ahlawat, Song Li, Eleni D. Pliakoni, Jeffrey Brecht, and Tie Liu. (2022) Identifying gene regulatory networks of senescence in postharvest broccoli (Brassica oleracea). Postharvest Biology and Technology. 183, 111729.
https://authors.elsevier.com/sd/article/S0925-5214(21)00268-4
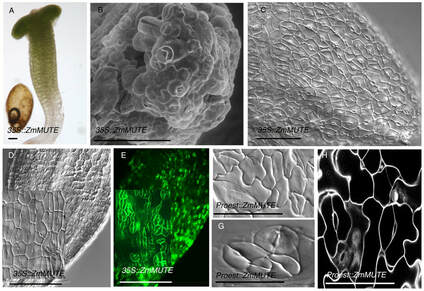
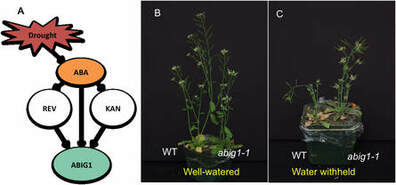
Jesus Preciado, Kevin Begcy, Liu T. (2022) The Arabidopsis ABIG1 is an Adaxial Regulator During Leaf Development. Journal of Experimental Botany. DOI: 10.1093/jxb/erab523
Yogesh Ahlawat, Tie Liu (2022) Varied expression of senescence-associated and ethylene-related genes during postharvest storage of Brassica vegetables. Int. J. Mol. Sci. 2021, 22(2), 839; https://doi.org/10.3390/ijms22020839.
2021
Taehoon Kim, Shafina Samraj, Juan Jimenez, Celina Gomez, Liu T, Kevin Begcy. (2021) Surveying the lettuce genome: Genome-wide identification of Hsfs and Hsps in response to different light conditions. BMC Plant Biology. 21, 185.
https://bmcplantbiol.biomedcentral.com/articles/10.1186/s12870-021-02959-x
Nidhi Sharma, Jesus Preciado, Tie Liu. (2021) KNOX HOMOLOGS SHOOT MERISTEMLESS (STM) and KNAT6 are epistatic to CLAVATA3 (CLV3) during embryonic meristem development in Arabidopsis thaliana. Molecular Biology Reports. 48(9):6291-6302. doi: 10.1007/s11033-021-066224.
|
|
Liu T, Talavera-Rauh F, Meghan Sharp, and Barton MK. (2022) Two AP2/ERF transcription factors are targets of SHOOTMERISTEMLESS (STM) gene that link abiotic stress and growth control. In submission.
|